

Sanjay J. Ingle*, K. K. Tapar
Vidyabharti College of Pharmacy, Amravati (MS), India
*Corresponding Author:
Sanjay J. Ingle
Vidyabharti College of Pharmacy, Amravati (MS), India
| Abstract |
Objective: The Quantitative structure-activity relationship (QSAR) analyses were carried out for a series of some 1,2-benzisoxazole derivatives to find out the structural requirements of their antipsychotic activities. Materials and Methods: Multiple linear regression (MLR) methodology coupled with various feature selection methods viz. stepwise (SW), genetic algorithm (GA) and simulated annealing (SA) were applied to develop QSAR models which were further validated for statistical significance and predictive ability by internal and external validation. Results: The statistically significant most excellent 2D QSAR model having correlation coefficient r2 = 0.7249 and cross validated squared correlation coefficient q2 = 0.6941with external predictive ability of pred_r2 = 0.6091was developed by SW-PLS with the descriptors like SaaCHE-index, SssOE-index, T_T_S_7, T_N_O_3 and SsssCHcount. Conclusions: The results of the present study may be helpful for the designing potent analogues as antipsychotic agents.
Keywords: QSAR, 1,2-benzisoxazole, antipsychotic agents, VLife MDS, MLR
| Introduction |
Psychoses is the mental disorder, with severe alteration of thought, behaviour, and capacity to identify reality and of awareness (delusions and hallucinations), affecting about 10% of world’s population. All clinically effective antipsychotics (except Clozapine like) have effective post synaptic dopaminergic D2 receptor blocking action and antipsychotic strength has shown well connection with their capacity to bind with D2 receptor (Lemke et al., 2008). Blockade action of dopamine in corpus striatum is responsible for the extrapyramidal symptoms (EPS) which is associated with antipsychotic drugs. In addition to blockade of dopaminergic receptor, some antipsychotics drugs like resperidone and clozapine also block 5HT system, which helps to lessen EP reactions, those related to their usefulness in improving negative symptoms (Nielsen and Nielsen, 2009). Some of antipsychotic drugs such as chlorpromazine have various adverse drug reactions such as parkinsonism, muscular rigidity, excessive salivation and many of the metabolic disturbances such as gynaecomastia, galactorrhea and aggravation of diabetes. Since epilepsy is very often associated with CNS psychiatric disorders, the drugs with antipsychotic as well as antiepileptic activity will be more beneficial. Further a need for more effective and less toxic antipsychotic drugs still exists. 1,2-benzisoxazole derivatives have reported consistent advances in the design of novel antipsychotic as well as anticonvulsant agents (Strupczewski et al. 1995, Uno et al., 1979).
Computational chemistry has developed as an important contributor to rational drug design. The Quantitative Structure-Activity Relationship (QSAR) move toward pioneered by Hansch et al. (2001) that helps to connect with definite biological actions or physical properties of large compounds with the measured or computed molecular properties, in terms of descriptors. QSAR methodologies save assets and expedite the process of the development of new drug molecules. There have been many QSAR researches related to design of antipsychotic drugs so far but a systematic QSAR study is yet to be carried out for series of 1,2-benzisoxazole derivatives.
The aim of present work is to derive some statistically significant QSAR models for 1,2-benzisoxazole derivatives for their antipsychotic activity and to relate antipsychotic activity to its physicochemical properties. The results obtained may contribute to further designing novel antipsychotic agents.
| Material and methods |
Data set
A data set of forty nine molecules from reported series of compounds 3-(1-substituted-4-piperidinyl)-1,2-benzisoxazoles for antipsychotic activities was taken for QSAR study (Strupczewski et al., 1995). Out of all these molecules, two molecules were discarded for which the precise data was not available (Table 1).
All 47 compounds were built on workspace of molecular modeling software VLife MDS 3.5 and then the structure was converted to three-dimensional space for further analysis. All these molecules were batch optimized for the reduction of energies using Merck molecular force field (MMFF) followed by considering distance-dependent dielectric constant of 1.0, convergence criterion or root-mean-square (RMS) gradient at 0.01 kcal/mol Å and the iteration limit to 10,000 (Halgren, 1996). The energy-minimized geometry was used for the calculation of the various 2D descriptors. The preprocessing of the independent variables (i.e., 2D descriptors) was done by removing invariable, which resulted in total 316 descriptors to be used for QSAR analysis. The sphere exclusion (SE) method (Golbraikh and Tropsha, 2003) was adopted for division of training and test data set comprising of 36 and 11 molecules, respectively, with dissimilarity value of 1.9 where the dissimilarity value gives the sphere exclusion radius. The unicolumn statistics of the training and test sets is reported in Table 2.
Feature selection and model development
Among several search algorithms, stepwise (SW) forward variable selection method (Darlington, 1990), genetic algorithms (GA) (Hasegawa, 1999) and simulated annealing (SA) (Zeng and Tropsha, 2000) based characteristic selection events are most popular for building QSAR models and can elucidate the situation more effectively.
To the selected equations, a cross-correlation limit was set at 0.5, the number of variables at 10, and the term selection criteria at q2. An F value was specified to evaluate the significance of a variable. The variance cutoff was set at auto scaling in which the number of random iterations was set at 100.
Model quality and validation
The developed QSAR models were evaluated using the following statistical measures: r2 (the squared correlation coefficient), Fischer’s value for statistical significance, q2 (correlation coefficient); pred_r2, r2 for external test set. The regression coefficient r2 is a relative measure of fit by the regression equation. It represents the deviation in the observed data that is explained by the regression. However, QSAR model is considered to be predictive, if the following conditions are satisfied: r2 > 0.6, q2 > 0.6 and pred_r2 > 0.5 (Golbraikh and Tropsha, 2002).
Internal validation was carried out using ‘leave-one-out’ (q2, LOO) method (Cramer, 1988). The cross-validated coefficient, q2, was calculated using the following equation:
where yi, and ŷi are the actual and predicted activity of the ith molecule in the training set, respectively, and ymean is the average activity of all molecules.
However, a high q2 value does not necessarily give a suitable representation of the real predictive power of the model. So, an external validation was also carried out in the present study. The external predictive power of the model was assessed by predicting pIC50 value of the 9 test set molecules, which were not included in the QSAR model development. The predictive ability of the selected model was also confirmed by pred_r2.
where yi, and ŷi are the actual and predicted activity of the ith molecule in the test set, respectively, and ymean is the average activity of all molecules in the training set.
| Results and discussion |
The QSAR study of 47 1,2-benzisoxazole derivatives for antipsychotic activity (Table 1) through MLR, PLS methodology, based on various selected methods viz. SW, GA and SA using VLife MDS 3.5 software that resulted following significant models, taking consider the term selection criterion as r2, q2 and pred_r2.
Table 1. Structures and antipsychotic activities of substituted 1,2- benzisoxazole derivatives
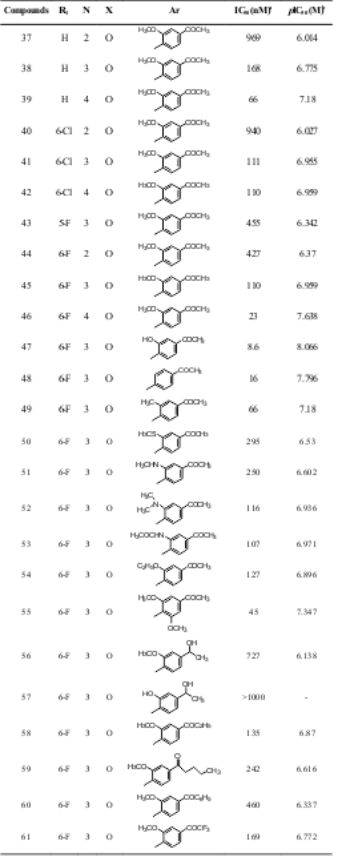
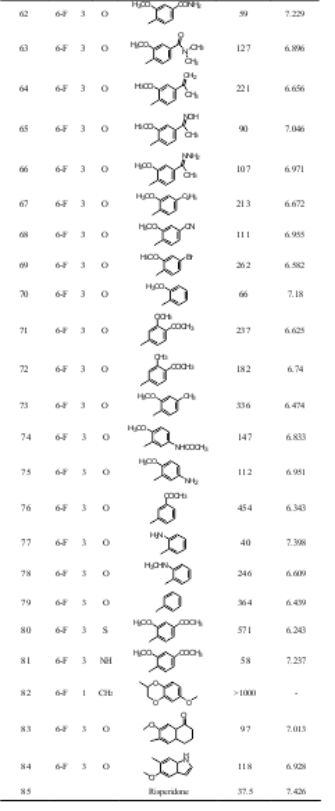 aIn-vitro antipsychotic activity against Dopamine receptor (D2); bpIC50 (M) = - log IC50 (M)
aIn-vitro antipsychotic activity against Dopamine receptor (D2); bpIC50 (M) = - log IC50 (M)
A Uni-Column statistics for training set and test set were generated to check correctness of selection criteria for trainings and test set molecules (Table 2).
This observation showed that test set was interpolative and derived within the minimum–maximum range of training set. The mean and standard deviation of pIC50 values of sets of training and test provide insights to relative difference of mean and point density distribution of two sets. Some statistically significant 2D QSAR models were chosen for discussion (Table 3).
Table 2. Uni-Column statistics of the training and test sets of substituted 1,2-benzisoxazole derivatives in 2D-QSAR
|
Data Set |
Sum |
Average |
Max |
Min |
Std Dev |
|
Training |
245.8590 |
6.8294 |
8.0660 |
6.0140 |
0.4260 |
|
Test |
75.8630 |
6.8966 |
7.7960 |
6.3420 |
0.4832 |
Max., maximum; Min., minimum; Std Dev, standard deviation
Table 3. Statistical results of some 2D-QSAR models generated for 3-(1-substituted-4-piperidinyl)-1,2-benzisoxazole derivative
|
S. No. |
Statistical parameters |
Model 1 (Best Model) |
Model 2 |
Model 3 |
|
1 |
r2 |
0.7249 |
0.7253 |
0.6294 |
|
2 |
q2 |
0.6941 |
0.5941 |
0.6486 |
|
3 |
pred_r2 |
0.6091 |
0.6530 |
0.5640 |
|
4 |
r2_se |
0.2327 |
0.2370 |
0.2619 |
|
5 |
q2_se |
0.2473 |
0.2866 |
0.2415 |
|
6 |
pred_r2se |
0.2669 |
0.2953 |
0.3015 |
|
7 |
F test |
15.1903 |
20.6393 |
15.8332 |
|
8 |
Zscore |
5.70240 |
4.82294 |
3.10712 |
|
9 |
ntraining |
36 |
36 |
36 |
Table 4. Inter-correlation matrix between descriptors used in best 2D-QSAR model 1
|
|
SaaCHE-index |
SssOE-index |
T_T_S_7 |
T_N_O_3 |
SsssCHcount |
|
SaaCHE-index |
1 |
|
|
|
|
|
SssOE-index |
0.094 |
1 |
|
|
|
|
T_T_S_7 |
-0.02 |
-0.239 |
1 |
|
|
|
T_N_O_3 |
0.087 |
-0.26 |
-0.103 |
1 |
|
|
SsssCHcount |
-0.05 |
0.174 |
-0.055 |
-0.127 |
1 |
Table 5. Comparative observed and predicted activities of 3-(1-substituted-4-piperidinyl)-1,2-benzisoxazole derivatives by 2D-QSAR models
*Experimental antipsychoic activity (pIC50) in molar concentration; aRes. = Exp. pIC50 – Pred. pIC50; #Compounds belong to test set
Model 1 (SW-PLS) (Best model)
The statistically best 2D-QSAR model (model 1, Eq. 3.9) using the SW-PLS method with r2 = 0.7249 was considered. The equation explains 72% (r2 = 0.72) of the total variance in the training set. It also has an internal (q2) and external (pred-r2) predictive ability of ~69% and ~61% respectively. The F-test = 15.19 shows the statistical significance of 99.99% of the model which means that probability of failure of the model is 1 in 10000.
The inter-correlation matrix between five descriptors with the biological activity for the best model 1 is given in Table 4 and contribution charts of models are shown in Figure 1.
The negative contribution of all above descriptors showed that decrease in the values of these descriptors would be beneficial for the antipsychotic activity of 3-(1-substituted-4-piperidinyl)-1,2-benzisoxazole derivatives. The residuals (difference between the actual and predicted activities) were found to be minimal and are presented in Table 5. Also the graphs for actual versus predicted activity for the series are shown in Figure 2 which shows good correlations.
| Conclusion |
The present work shows about a set of compounds with antipsychotic activities of 1,2-benzisoxazoles treated statistically to uncover the molecular characteristics which are important to high activity. The developed models were analyzed and validated for their statistical significance and external prediction power. The awareness and understanding of the descriptors involved in the activity of these compounds could provide a great opportunity for the ligand structures design with appropriate features. These theories, by which these features affect the biological activity, by binding to the respective receptor target. The results derived may be useful in further designing more novel antipsychotic agents in series.
| References |
Clark M, Cramer RD. 1989. III, Van, O.N. Validation of the general purpose Tripose 5.2 force field. Journal of Computational Chemistry, 10: 982–1012.
Darlington RB. 1990. Regression and Linear Models, McGraw-Hill, New York.
Golbraikh A, Tropsha A. 2002. Beware of q2!. Journal of Molecular Graphics and Modelling, 20: 269-276.
Golbraikh A, Tropsha A. 2003. QSAR modeling using chirality descriptors derived from molecular topology. Journal of Chemical Information and Modeling, 43: 144–154.
Halgren TA. 1996. Merck molecular force field. III. Molecular geometries and vibrational frequencies for MMFF94Journal of Computational Chemistry, 17: 553–586.
Hansch C, Leo A. 1995. Exploring QSAR. In Fundamentals and Applications in Chemistry and Biology; Heller, S.R., Ed.; American Chemical Society: Washington.
Hasegawa K, Kimura T, Funatsu K. 1999. GA strategy for variable selection in QSAR studies: Enhancement of comparative molecular binding energy analysis by GA-based PLS method. Quantitative Structure-Activity Relationships, 18: 262-272.
Lemke T L, Williams DA, Roche VF, Zito SW. 2008. Foye's Principles of Medicinal Chemistry, 6th ed.; Lippincott Williams & Wilkins, pp 601-627.
Nielsen RE, Nielsen J. 2009. Antipsychotic drug treatment for patients with schizophrenia: theoretical background, clinical considerations and patient preferences. Clinical Medicine Insights: Therapeutics, 1: 1053–1068.
Strupczewski JT, Bordeau KJ, Chiang Y, Glamkowski EJ, Conway PG, Corbett R, Hartman HB, Szewczak MR, Wilmot CA, Helsley GC. 1995. 3-[[(Aryloxy)alkyl]piperidinyl]-1,2-benzisoxazoles as D2/5-HT2 antagonists with potential atypical antipsychotic activity: antipsychotic profile of iloperidone (HP 873). Journal of Medicinal Chemistry, 38(7):1119-31.
Uno H, Kurokawa M, Masuda Y, Nishimura H. 1979. Studies on 3-substituted 1,2-benzisoxazole derivatives and their anticonvulsant activities . Journal of Medicinal Chemistry, 22: 180-183.
VLifeMDS3.5 Molecular Design Suite, Vlife Sciences Technologies Pvt. Ltd. Pun, India (www.vlifesciences.com) (2008).
Zheng W, Tropsha A. 2000. Novel variable selection quantitative structure–property relationship approach based on the k-nearest neighbor principle. Journal of Chemical Information and Computer Sciences, 40: 185-194.